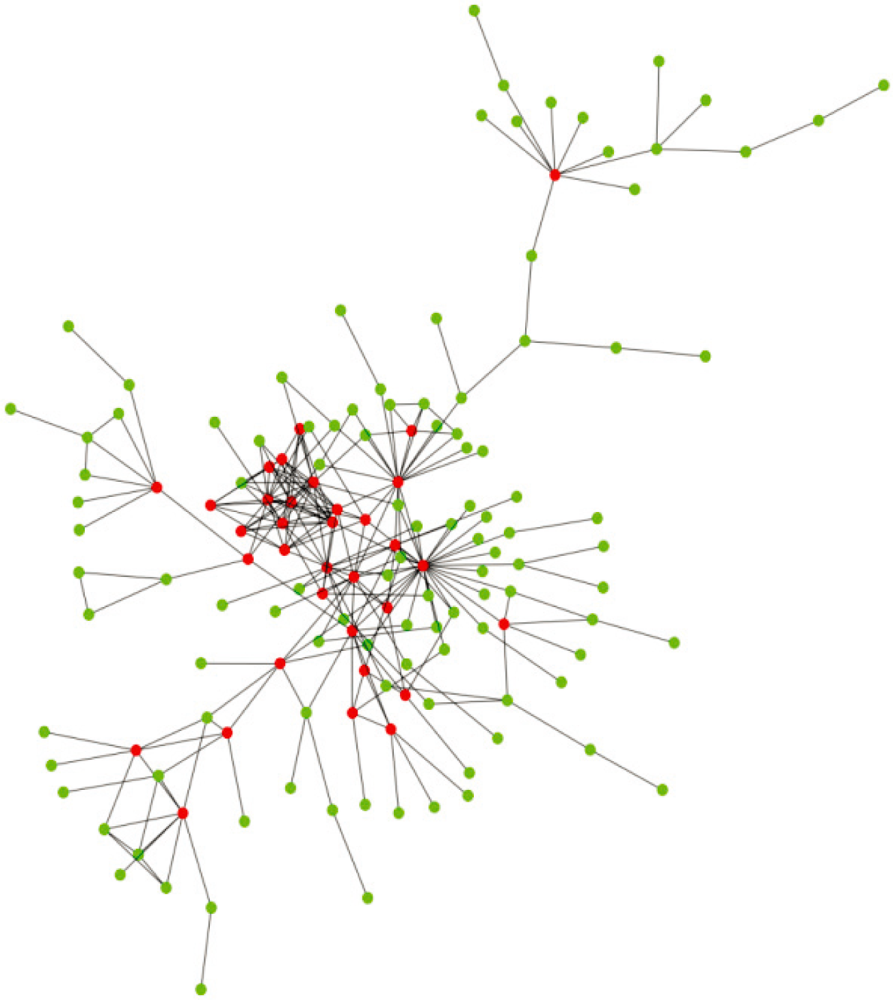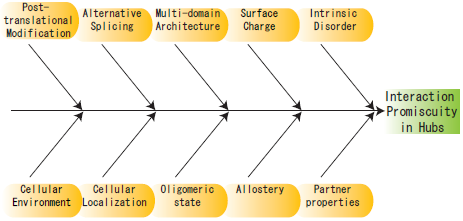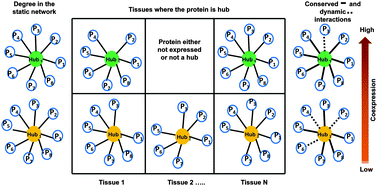
Interaction and localization diversities of global and local hubs in human protein–protein interaction networks - Molecular BioSystems (RSC Publishing)

Protein–protein interaction networks: how can a hub protein bind so many different partners?: Trends in Biochemical Sciences
![Ageing- and AAA-associated differentially expressed proteins identified by proteomic analysis in mice [PeerJ] Ageing- and AAA-associated differentially expressed proteins identified by proteomic analysis in mice [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/13129/1/fig-5-full.png)
Ageing- and AAA-associated differentially expressed proteins identified by proteomic analysis in mice [PeerJ]
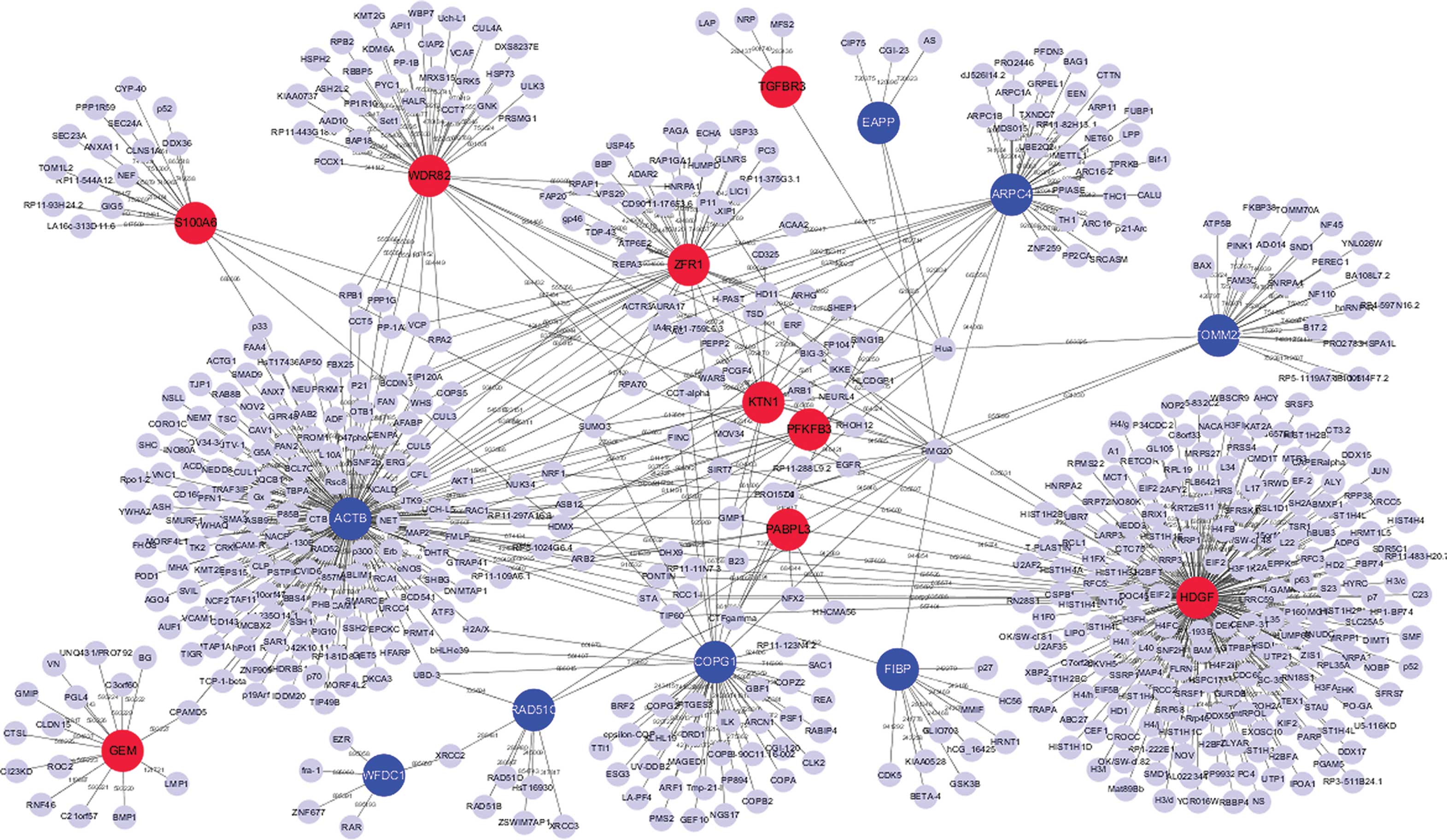
Integrated analysis of differential gene expression profiles in hippocampi to identify candidate genes involved in Alzheimer's disease

Figure 4 | Computational Simulations to Predict Creatine Kinase-Associated Factors: Protein-Protein Interaction Studies of Brain and Muscle Types of Creatine Kinases

Protein-protein interactions network model underlines a link between hormonal and neurological disorders - ScienceDirect
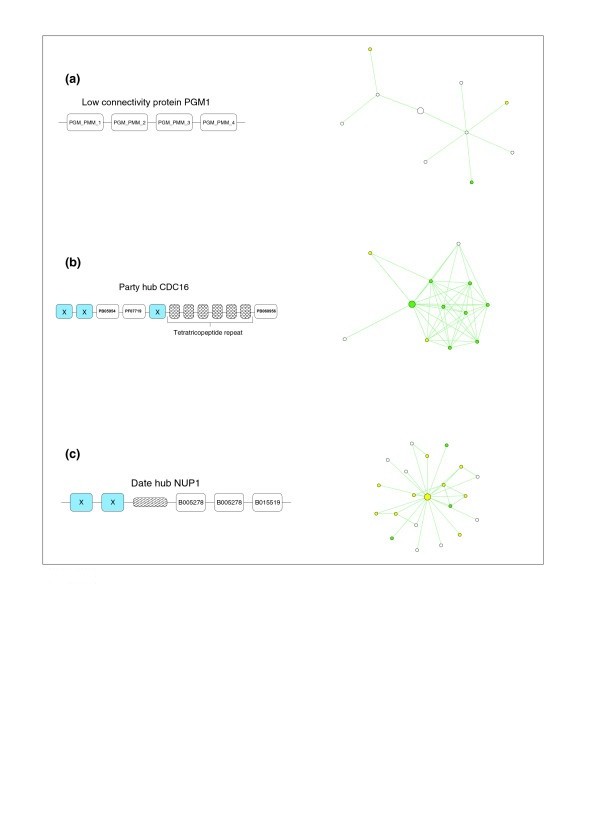

What properties characterize the hub proteins of the protein-protein interaction network of Saccharomyces cerevisiae? | Genome Biology | Full Text

Figure 1 | Computational Simulations to Predict Creatine Kinase-Associated Factors: Protein-Protein Interaction Studies of Brain and Muscle Types of Creatine Kinases

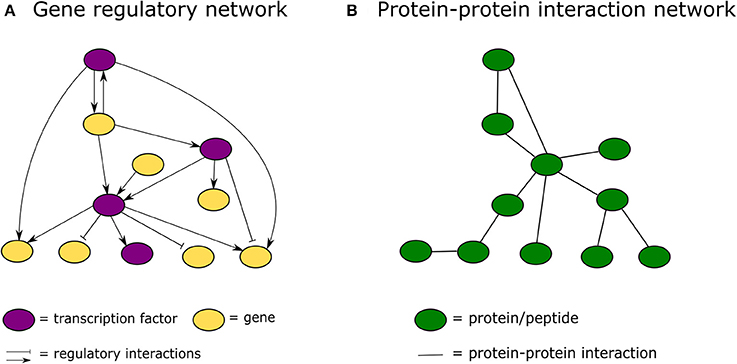
Protein-protein interaction network and molecular hubs. (A) Schematic... | Download Scientific Diagram
Predicting the Binding Patterns of Hub Proteins: A Study Using Yeast Protein Interaction Networks | PLOS ONE

Protein-protein interaction (PPI) network analysis reveals important hub proteins and sub-network modules for root development in rice (Oryza sativa) | bioRxiv
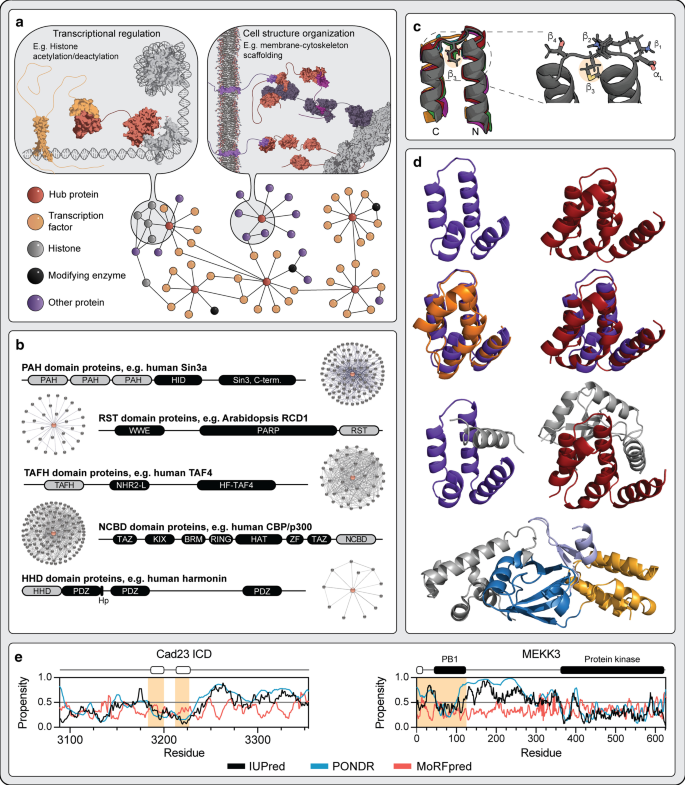
Connecting the αα-hubs: same fold, disordered ligands, new functions | Cell Communication and Signaling | Full Text